Imaging aggregation of histologically stained F-actin ex vivo using the contrast in Förster resonance energy transfer obtained from 2D polarization fluorescence imaging (2D-POLIM)
- Abstract number
- 6
- Presentation Form
- Oral
- DOI
- 10.22443/rms.elmi2021.6
- Corresponding Email
- [email protected]
- Session
- Live and Functional Imaging Technologies Part 1
- Authors
- Daniela Täuber (3, 5), Juanzi Shi (1), Petra Babic (2), Kay-Jovanna Benecke (2), Michael Bauer (2), Rainer Heintzmann (3, 5, 6), Ivan G. Scheblykin (1), Adrian T. Press (2, 4)
- Affiliations
-
1. Chemical Physics, Lund University, Lund, Sweden
2. Department of Anesthesiology and Intensive Care Medicine, Jena University Hospital, Jena, Germany
3. Leibniz Institute of Photonic Technology, Albert-Einstein Str. 9, 07745 Jena, Germany
4. Friedrich-Schiller University, Faculty of Medicine, Jena Germany
5. Institute of Physical Chemistry, Friedrich-Schiller University, Jena, Germany
6. Abbe Center of Photonics, Friedrich-Schiller University, Jena, Germany
- Keywords
- fluorescence polarization imaging
- phalloidin staining
- severe sepsis
- liver tissue ex vivo
- polymicrobial infection
- Abstract text
Summary
2D polarization fluorescence imaging (2D-POLIM) was applied to phalloidin-DY490 stained mouse liver tissue of mice from different treatment regimes of AS605240 and respective vehicle groups in the context of systemic infection. The obtained contrasts in excitation energy transfer showed higher values for mice with pathogenic conditions than for the healthy control and for those treatment regimes that were found to improve the health condition.
Introduction
Polarization resolved fluorescence microscopy provides access to determine the intermolecular distance between similar fluorescence labels via polarization resolved evaluation of Förster resonance energy transfer (homo-FRET, emFRET). In general, FRET can be used to discriminate small aggregates of fluorescent molecules from just high concentration without aggregation. Polarization resolved evaluation of FRET circumvents the need for a pair of different fluorescence labels required for spectrally resolved FRET measurements. However, the application of conventional fluorescence anisotropy measurements to anisotropic tissue and cell samples may show a dependence on the lab frame. This is avoided by 2D-POLIM, which provides the full in-plane information of the polarization state of the sample by synchronized control of the orientation angles of linearly polarized excitation and of fluorescence detection.[1] 2D-POLIM was successfully applied to investigate the early aggregation of GFP-labeled human α-synuclein in models of Parkinson’s disease ex vivo.[2]
Histological staining provides access to the investigation of cells and tissue ex vivo without the need for genetic modifications. Phalloidin complexes bind to Fibrillary (F)-Actin fibers in a defined geometry, and are thus suitable as histological stains for fluorescence polarization microscopy.[3] In polymicrobial sepsis the microvilli containing brush borders in hepatocytes are lost,[4] which is characterized by changes in the condensation of F-Actin. We applied 2D-POLIM to phalloidin stained healthy and pathologic HepaRG cells and liver tissue to investigate the particular aggregation of F-Actin.
Methods/Materials
2D-POLIM was performed on snap frozen, freshly formalin fixed liver tissue stained with DY-490 conjugated Phalloidin (Dyomics GmbH). The imaging setup and image analysis was recently published for biological samples and used with 488 nm excitation.[2] In brief, 2D-POLIM is based on a wide-field fluorescence microscope where the sample is illuminated with light linearly polarized within a plane oriented at φex. The fluorescence signal is imaged on a camera via a polarization analyzer oriented at φem. Both angles (φex and φem) are varied during imaging of the fluorescence intensity I. The obtained polarization portrait I(φex, φem) is analyzed using a model for energy transfer which provides a normalized parameter ε quantifying the energy transfer efficiency between fluorescent labels.[1,2]
The liver tissue was obtained from mice in a model of peritoneal sepsis with different treatment regimens of the PI3K-γ inhibitor AS605240 (AS), including a liposome carrier for its targeted release, and respective vehicle groups.[5,6] HepaRG cells treated with a cytokine mix were used for comparison, as described elsewhere.[5]
Results and Discussion
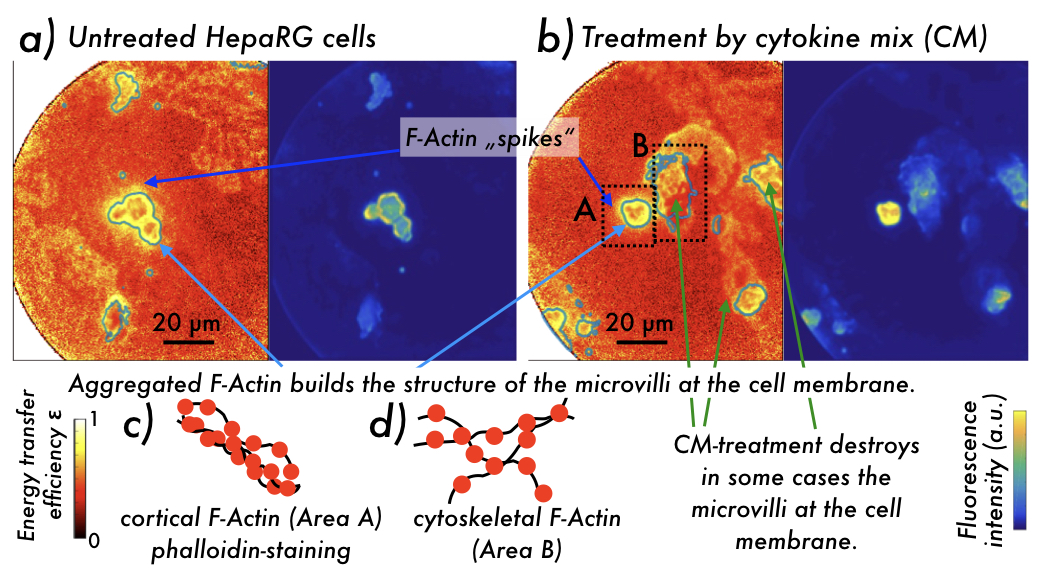
Figure 1: Imaging contrast in (left) energy transfer efficiency (homo-FRET metrics) ε and (right) fluorescence intensity obtained using 2D POLIM from phalloidin-DY490 stained HepaRG cells which were a) untreated, and b) treated by a cytokine mix (CM). Greenish contour lines in the ε-contrast images highlight areas of fluorescence intensity above a threshold. Highly aggregated F-Actin is seen at intact cell membranes building bile canaliculi, and in spikes extending into the cells. c,d) Schemes showing c) highly aggregated cortical, and d) less aggregated cytoskeletal F-Actin stained by a phalloidin-DY490 complex, resulting in the structures seen in b) area A and B, respectively.
2D-POLIM applied to stained HepaRG cells showed high energy transfer efficiency ε in areas, where these cells form apical canalicular membrane-structures.
Further areas showing slightly less high ε can be assigned to less aggregated cytoskeletal F-Actin, see Figure 1.
2D-POLIM applied to liver tissue from differently treated mice revealed significant differences in the molecular arrangement of F-Actin. In animals suffering polymicrobial sepsis the aggregation of F-Actin was significantly increased. Treatment with AS and with AS loaded liposomes (T-LipoAS) 6 h post infection and also in healthy controls (sham) decreased the aggregation of F-Actin. The observation agrees with a higher probability of survival for animals treated by AS605240.[6] These findings from a first application to histologically stained cells and tissue ex vivo are very promising for designing further studies using 2D-POLIM to enhance the understanding of pathways leading to organ failure in polymicrobial sepsis.
Conclusion
Significant differences in the molecular arrangement of F-Actin were found in liver tissue from animals in healthy and pathological conditions. The observations of increased and decreased aggregation of F-Actin are in accordance with the probability of survival for the respective animal groups. These findings approve the successful application of 2D-POLIM to histologically stained liver tissue and cells.
- References
[1] R. Camacho, D. Täuber, I. G. Scheblykin, Adv. Mater. 2019, 31, 1805671.
[2] R. Camacho, D. Täuber, C. Hansen, J. Shi, L. Bousset, R. Melki, J.-Y. Li, I. G. Scheblykin, Commun. Biol. 2018, 1, 157.
[3] P. Nordenfelt, T. I. Moore, S. B. Mehta, J. M. Kalappurakkal, V. Swaminathan, N. Koga, T. J. Lambert, D. Baker, J. C. Waters, R. Oldenbourg, T. Tani, S. Mayor, C. M. Waterman, T. A. Springer, Nat. Commun. 2017, 8, 2047.
[4] P. Recknagel, F. A. Gonnert, M. Westermann, S. Lambeck, A. Lupp, A. Rudiger, A. Dyson, J. E. Carré, A. Kortgen, C. Krafft, J. Popp, C. Sponholz, V. Fuhrmann, I. Hilger, R. A. Claus, N. C. Riedemann, R. Wetzker, M. Singer, M. Trauner, M. Bauer, PLoS Med. 2012, 9, e1001338.
[5] P. Martinac, A. T. Press, A. Medyukhina, K.-J. Benecke, J. Shi, D. Täuber, S. Hoeppener, Z. Cseresnyes, I. G. Scheblykin, M. H. Gräler, I. Rubio, M. T. Figge, U. S. Schubert, M. Bauer, Infection 2019, 47, S6.
[6] A. T. Adrian Press, P. Babic, B. Hoffmann, T. Müller, W. Foo, W. Hauswald, J. Benecke, M. Beretta, Z. Cseresnyés, S. Hoeppener, I. Nischang, S. M. Coldewey, M. H. Gräler, R. Bauer, F. Gonnert, N. Gaßler, R. Wetzker, M. Thilo Figge, U. S. Schubert, M. Bauer, bioRxiv 2021, 2021.01.20.427305.